Improving the efficiency of process development for biopharmaceutical manufacturing methods, e.g. recombinant protein production requires genome wide insight into fundamental meatabolic and regulatory mechanisms of the expression system for the achievement of further progress in process development. Recently developed new tools for micro-biochemical analysis of DNA and proteins in combination with high troughput platforms, such as DNA microarrays and differential 2-D electrophoresis create new analytical opportunities for large scale acquisition of whole genome and proteome data.
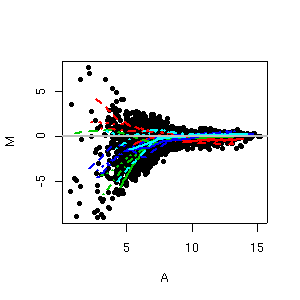
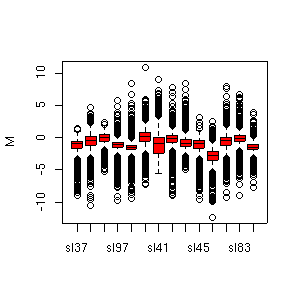
Microarrays are tools to study the expression level of thousands of individual DNA sequences simultaneously. As researchers have to manage massive amouts of data, statistical considerations become more and more important in the analysis of microarrays. Careful statistical design and analysis are essential to improve the efficiency and reliability of microarray experiments as they generate large and complex multivariate datasets. Access to an efficient statistical computing environment is essential for the analysis of gene expression data. Such an environment is provided in R through the numerous extension packages developed by the Bioconductor project. Hybridisation ratios are analyzed using Bioconductor package limma. Data analysis includes data display and exploration, as well as normalization, differential expression analysis and clustering methods.